If you’ve ever worked with Python for data science, machine learning, or scientific computing, you’ve probably heard this sentence:
“Use NumPy — it’s much faster than plain Python.”
But why is NumPy fast?
What really happens under the hood?
And how does its C backend make such a massive difference?
Let’s break it down in simple terms.
The Problem with Pure Python Loops
Python is an interpreted, high-level language. That makes it easy to read and write, but not always fast.
When you write a loop like this:
for i in range(len(a)):
c[i] = a[i] + b[i]
Python does a lot of work behind the scenes:
- Type checking on every iteration
- Bounds checking
- Dynamic memory handling
- Function calls for even basic operations
This overhead happens millions of times in large datasets — and that’s where performance suffers.
NumPy Solves This with C Under the Hood
NumPy is written mostly in C, not Python.
When you call a NumPy operation like:
c = a + b
You are not looping in Python.
Instead:
- Python hands control to NumPy
- NumPy executes a compiled C loop
- The loop runs directly on raw memory
- The result is returned to Python
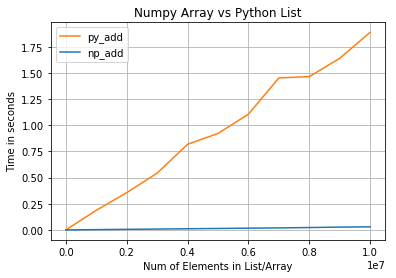
This is why NumPy operations are often 10x to 100x faster than equivalent Python code.
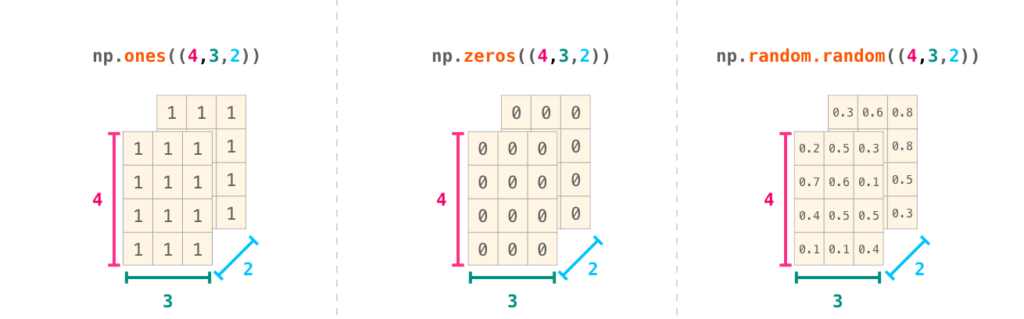
Contiguous Memory: The Secret Weapon
NumPy arrays store data in contiguous blocks of memory, just like C arrays.
This gives multiple advantages:
- Better CPU cache usage
- Fewer memory lookups
- Faster sequential access
In contrast, Python lists store references to objects, scattered across memory, which slows things down.
Because NumPy knows:
- The data type
- The size of each element
- The exact memory layout
It can perform operations extremely efficiently.
Vectorization: No Python Loops
One of NumPy’s biggest performance benefits is vectorization.
Vectorization means:
- Operations run on entire arrays at once
- No explicit loops in Python
- Computation happens in optimized C code
Example:
a = a * 2
This single line replaces:
- A Python loop
- Multiple function calls
- Repeated type checks
And executes as a tight, low-level C loop instead.
Optimized Libraries Behind NumPy
NumPy doesn’t just use C — it also relies on highly optimized native libraries, such as:
- BLAS (Basic Linear Algebra Subprograms)
- LAPACK
- Intel MKL (on some systems)
These libraries are:
- Written in C and Fortran
- Tuned for specific CPUs
- Optimized with SIMD instructions and multi-threading
That’s why matrix multiplication, linear algebra, and numerical operations are blazing fast in NumPy.
Fixed Data Types Reduce Overhead
NumPy arrays have a fixed data type (int32, float64, etc.).
This means:
- No dynamic type checking
- No resizing during computation
- Predictable memory usage
Python lists, on the other hand, can mix types, which forces Python to do extra checks every time.
Why NumPy Is Essential for Data Science
NumPy’s speed makes it the foundation for:
- Pandas
- SciPy
- Scikit-learn
- TensorFlow
- PyTorch (internally)
Without NumPy’s C backend, modern data science and machine learning in Python would simply not be practical.
Final Thoughts
NumPy is fast not because Python is fast, but because Python gets out of the way.
By:
- Offloading heavy computation to C
- Using contiguous memory
- Eliminating Python loops
- Leveraging optimized math libraries
NumPy delivers near low-level performance with high-level simplicity.
That’s the real magic of NumPy — Python syntax with C-level speed.